NEXT GENERATION SEQUENCING
In our search for the right bacteria, we optimised a Next Generation Sequencing platform for microbiome analysis. But what exactly is NGS? And what can you do with it?
WHAT IS NEXT GENERATION SEQUENCING?
In the past, people determined the DNA code gene by gene. Fortunately, the techniques for analysing genetic material such as DNA and RNA have greatly improved over the last 20 years. Think of the Covid tests in which a PCR (Polymerase Chain Reaction) technique is used to quantify the amount of viral genetic material, RNA in the case of the SARS-CoV2 virus. This analysis is based specifically on the presence of a short genetic sequence which is then amplified.
Next generation sequencing (NGS) goes one step further and does not only look at a specific sequence but aims to read the code of all DNA or RNA present in a certain sample. The Human Genome Project in particular, sponsored by the National Institute of Health (NIH), has provided an enormous boost in the improvement of NGS techniques. After obtaining an enormous amount of data (large text files with the base pair code of DNA; i.e. ACTGATTTCCC...), bioinformatics is used to find where these codes correspond to. Depending on the sample and the research question, one can specifically look for similarities with the genetic codes of plants, animals, people, viruses, fungi, bacteria,... or all of these at the same time.
WHAT CAN YOU
DO WITH IT?
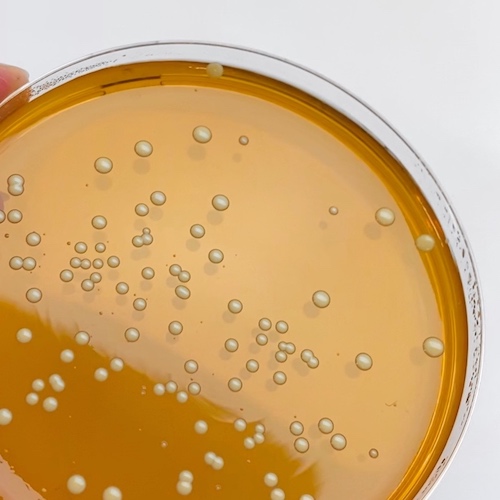
First we take a sample, for example using a swab of the skin, and we can purify the DNA from the sample. With the help of NGS, all DNA material is read out. Then we can find out which bacteria are present on the skin. Because of the enormous amount of information, we not only know whether certain Staphylococcus spp. are present, but we can even check whether a specific strain such as Staphylococcus aureus MRSA, the well-known hospital bacteria, is present.
Of course, this is only one specific example; the possibilities and techniques are continuously being improved and expanded. It gives us and other researchers around the world a powerful toolbox to gain more insights into various complex research domains!

In collaboration with the University of Antwerp (Laboratory of Applied Microbiology and Biotechnology - Prof. Sarah Lebeer), we optimised a NEXT-GEN-SEQUENCING platform for microbiome analysis. This allowed us to analyse 16S rRNA (bacteria) and ITS (fungi) for low biomass samples (such as the skin).
Proudly mentioned in







Menu
Contact
+32 (0)3 443 04 70
info@yun.be
Galileilaan 15
2845, Niel
Belgium
Social
With the support of